Huixia Lu, a scholarship student from the China Scholarship Council (2016), defends her thesis on the behavior of proteins and small molecules in the environment of cell membranes
Oct 05, 2020
La Huixia Lu defended his thesis directed by Jordi Martí Rabassa. on September 29, 2020 at the Nort Campus of the UPC, entitled "Exploring free-energy landscapes and microscopic interactions of selected small-molecules and proteins with cell membranes", the thesis describes the interactions at the atomic level between molecular systems of biological interest and pharmacist with the constitutive lipids of cell membranes.
The present Thesis is devoted to the study of the physical-chemical properties of selected small-molecules (such as amino-acids like tryptophan or hormones like melatonin) and proteins (such as KRAS-4B) absorbed in model lipid bilayers located at physiological environments. Since in such conditions biological membranes composed of phospholipids and cholesterol are surrounded by electrolyte solutions, understanding the interactions of the small molecule or protein with the surrounding phospholipids, cholesterol, water and all sorts of ion species is a topic of great fundamental importance. In particular, the present Thesis has advanced into the analysis of the structural and energetic aspects of an oncogenic protein from the Ras family, characterising the physical conditions that allow such protein to remain anchored to the cell. The findings reported in the Thesis may help to shed light in the understanding of a wide variety of cancers, with direct impact on the design of drugs or treatments useful for curation.
The lipids considered in this Thesis include the saturated lipids dimyristoilphosphatidylcholine and dipalmytoilphosphatidylcholine, the unsaturated lipids dioleoylphosphatidylcholine and dioleoylphosphatidylserine and cholesterol. Classical molecular dynamics simulations and well-tempered metadynamics simulations have been applied in this thesis so that all considered systems have been modelled and simulated at the all-atom level, with systems containing up to 200000 atoms.
Using classical molecular dynamics simulations at the microsecond time scale, we studied the microscopic structure and dynamics of the small-molecules and KRas-4B proteins, the latter in the wild-type and mutated (oncogenic) forms. The cell membrane has been always considered in the liquid crystalline phase, what in some cases required to rise the temperature of the system up to 323 K. Structural properties such as the area per lipid and thickness of the membrane, density profiles, deuterium-order parameters, orientational distributions and the extent of water penetration in the membrane have been analysed. Molecular self-diffusion and spectral densities of atomic species reveal a variety of time scales playing a role in membrane dynamics. The physical meaning of all spectral features from lipid atomic sites is analysed and correlated with experimental data. Most relevant have been the location of individual sites of binding of probes at the interface of the membrane. Finally, using reversible work techniques, we estimated the extent of free energy required to form such liaisons.
By applying 1-microsecond well-tempered metadynamics simulations, we have performed systematic free energy calculations of probe binding to the membrane and water for the first time. Free energy landscapes unveil specific binding behaviour of small-molecules and proteins at phospholipid membranes. This Thesis provides a general methodology to explore such free energy landscapes at complex biological interfaces which can be extended to study other interactions of interest between molecules, peptides, proteins or drugs and charged head-groups in colloidal chemistry and biology.
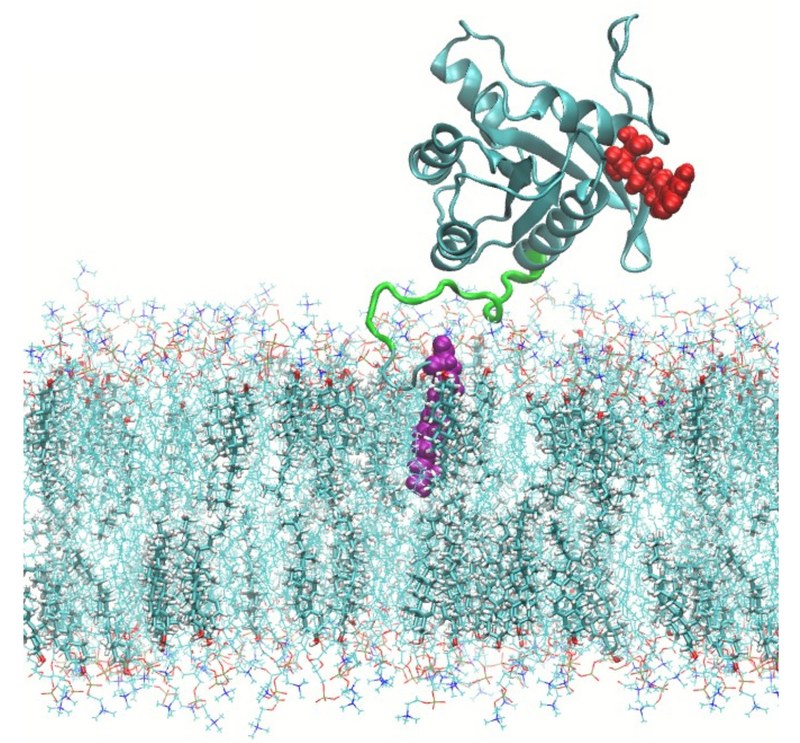
We further applied this methodology to study the case of a prototypical oncogenic protein (KRas), being able to produce a wide variety of cancers. Our results from resulting free energy landscapes indicate the existence of specific hydrogen-bonding connections between parts of the protein (hypervariable region and farnesylated tail) that might be responsible of the permanent infection of healthy cells through its anchoring at the interface of the membrane.
Share: